array of ai 3d drawing nature geometry
Abstract
Constructive molecular representation learning is of great importance to facilitate molecular property prediction. Recent advances for molecular representation learning have shown great promise in applying graph neural networks to model molecules. Moreover, a few recent studies pattern self-supervised learning methods for molecular representation to address insufficient labelled molecules; however, these self-supervised frameworks treat the molecules every bit topological graphs without fully utilizing the molecular geometry data. The molecular geometry, also known as the three-dimensional spatial construction of a molecule, is critical for determining molecular properties. To this end, we advise a novel geometry-enhanced molecular representation learning method (GEM). The proposed Gem has a specially designed geometry-based graph neural network architecture as well equally several dedicated geometry-level self-supervised learning strategies to acquire the molecular geometry knowledge. We compare Gem with various land-of-the-art baselines on different benchmarks and show that information technology tin considerably outperform them all, demonstrating the superiority of the proposed method.
Principal
Molecular property prediction has been widely considered as ane of the nearly critical tasks in computational drug and materials discovery, equally many methods rely on predicted molecular properties to evaluate, select and generate moleculesane,2. With the development of deep neural networks (DNNs), molecular representation learning exhibits a keen reward over characteristic engineering-based methods, which has attracted increasing enquiry attending to tackle the molecular belongings prediction trouble.
Graph neural networks (GNNs) for molecular representation learning have recently become an emerging research surface area, which regard the topology of atoms and bonds as a graph, and propagate messages of each element to its neighboursthree,4,5,6. However, one major obstacle to hinder the successful awarding of GNNs (and DNNs) in molecule property prediction is the scarity of labelled data, which is too a common research challenge in natural language processingvii,8 and computer visionix,ten communities. Inspired by the success of cocky-supervised learning, recent studiesiv,xi start to use large-scale unlabelled molecules in a cocky-supervised methodology to pre-train the molecular representation and and then use a small number of labelled molecules to fine melody the models, achieving substantial improvements.
Existing self-supervised learning techniques for GNNs4,xi only consider the topology information of the molecules, neglecting the molecular geometry, that is, the three-dimensional spatial structure of a molecule. These works conduct self-supervised learning by masking and predicting in nodes, edges or contexts in the topology4,11. Yet these tasks only enable the model to acquire the laws of molecular graph such every bit which cantlet/group could exist connected to a double bail, and lack the ability to learn the molecular geometry noesis, which plays an important role in determining molecules' concrete, chemical and biological activities. For example, the water solubility (a critical metric of drug-likeness) of the two molecules illustrated in Fig. 1 is dissimilar due to their differing geometries, even though they have the same topology. Cis-platin and trans-platin are another case of molecules with the same topology but different geometries: cis-platin is a pop chemotherapy drug used to treat a number of cancers, whereas trans-platin has no cytotoxic activity12.
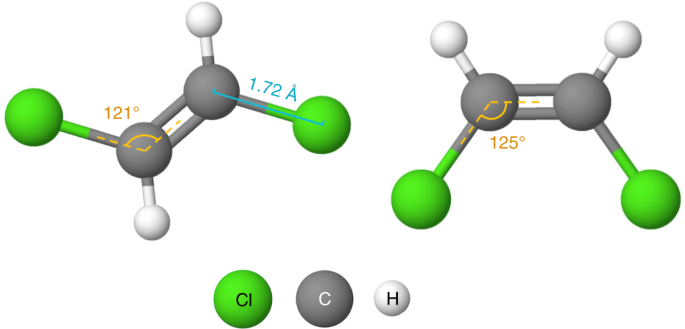
The two chlorine atoms are on different sides in trans-1,two-dichloroethene (left) but the same side in cis-1,2-dichloroethene (right).
Although incorporating geometric data into graph architectures to benefit some molecular property estimation tasks has attracted research attention in contempo years13,fourteen,xv,16,17, there is still a need to utilize the molecular geometry data to develop a cocky-supervised learning image for property prediction. We argue that adopting the self-supervised learning to estimate the geometry can contribute to facilitating the model'southward chapters in predicting various backdrop. Self-supervised learning can take advantage of the big-scale unlabelled molecules with coarse 3-dimensional spatial structures to amend learn the molecular representation, where the coarse three-dimensional spatial structures can be efficiently calculated by cheminformatics tools such as RDKit (https://www.rdkit.org/). Past geometry-level self-supervised learning, the pre-trained model is capable of inferring the molecular geometry past itself.
To this end, we propose a novel geometry-enhanced molecular representation learning method (GEM). Showtime, to make the message passing sensitive to geometries, we model the effects of atoms, bonds and bond angles simultaneously by designing a geometry-based GNN compages (GeoGNN). The compages consists of two graphs: the showtime graph regards the atoms as nodes and the bonds as edges, whereas the second graph regards the bonds equally nodes and the bond angles as edges. Second, we pre-railroad train the GeoGNN to learn the chemic laws and the geometries from big-scale molecules with coarse three-dimensional spatial structures, designing diverse geometry-level self-supervised learning tasks. To verify the effectiveness of the proposed GEM, we compared it with several state-of-the-fine art (SOTA) baselines on 15 molecular property prediction benchmarks, among which Gem achieves 14 SOTA results.
Our contributions can be summarized as follows:
-
We propose a novel geometry-based GNN to encode both the topology and geometry information of molecules.
-
We pattern multiple geometry-level self-supervised learning tasks to learn the molecular spatial knowledge from large-calibration molecules with fibroid spatial structures.
-
We evaluated GEM thoroughly on various molecular belongings prediction datasets. Experimental results demonstrate that Jewel considerably outperforms competitive baselines on multiple benchmarks.
Preliminaries
Graph-based molecular representation
A molecule consists of atoms and the neighbouring atoms are connected by chemical bonds, which tin be represented by a graph \(G=({{{\mathcal{V}}}},{{{\mathcal{E}}}})\), where \({{{\mathcal{V}}}}\) is a node ready and \({{{\mathcal{Eastward}}}}\) is an edge prepare. An atom in the molecule is regarded equally a node \(v\in {{{\mathcal{5}}}}\) and a chemic bail in the molecule is regarded every bit an edge \((u,5)\in {{{\mathcal{East}}}}\) connecting atoms u and v.
Graph neural networks are message-passing neural networks18, making them useful for predicting molecular properties. Post-obit the definitions of the previous GNNs19, the features of a node five are represented by 10 v and the features of an border (u,v) are represented by x u v . Taking node features, border features and the graph structure as inputs, a GNN learns the representation vectors of the nodes, where the representation vector of a node v is denoted by h v . A GNN iteratively updates a node's representation vector past aggregating the messages from the node'southward neighbours. Finally, the representation vector h G of the entire graph can be obtained by pooling over the representation vectors {h five } of all the nodes at the last iteration. The representation vector of the graph h G is utilized to estimate the molecular backdrop.
Pre-training methods for GNNs
In the molecular representation learning community, recently several works4,11,20 have explored the ability of self-supervised learning to improve the generalization ability of GNN models on downstream tasks. They mainly focus on two kinds of self-supervised learning tasks: the node-level (edge-level) tasks and the graph-level tasks.
The node-level self-supervised learning tasks are devised to capture the local domain knowledge. For example, some studies randomly mask a portion of nodes or sub-graphs and so predict their backdrop by the node/edge representation. The graph-level self-supervised learning tasks are used to capture the global information, like predicting the graph properties by the graph representation. Usually, the graph properties are domain-specific cognition, such as experimental results from biochemical assays or the being of molecular functional groups.
The Jewel framework
This department introduces the details of our proposed geometry-enhanced molecular representation learning method (GEM), which includes ii parts: a novel geometry-based GNN and various geometry-level self-supervised learning tasks.
GeoGNN
Nosotros propose a GeoGNN that encodes molecular geometries past modelling the cantlet–bond–angle relations, distinguishing them from traditional GNNs, which but consider the relationship between atoms and bonds.
For a molecule, we denote the atom set every bit \({{{\mathcal{V}}}}\), the bond set every bit \({{{\mathcal{E}}}}\), and the bail angle ready as \({{{\mathcal{A}}}}\). We introduce atom–bail graph Chiliad and bond–bending graph H for each molecule, equally illustrated in Fig. 2a. The cantlet–bond graph is divers every bit \(G=({{{\mathcal{V}}}},{{{\mathcal{E}}}})\), where cantlet \(u\in {{{\mathcal{V}}}}\) is regarded as the node of 1000 and bond \((u,v)\in {{{\mathcal{E}}}}\) as the edge of G, connecting atoms u and v. Similarly, the bond–bending graph is defined as \(H=({{{\mathcal{East}}}},{{{\mathcal{A}}}})\), where bond \((u,v)\in {{{\mathcal{East}}}}\) is regarded as the node of H and bail angle \((u,v,west)\in {{{\mathcal{A}}}}\) every bit the edge of H, connecting bonds (u,v) and (five,w). We use x u as the initial features of cantlet u, ten u v as the initial features of bond (u,5), and x u v due west as the initial features of bail bending (u,v,w). The atom–bond graph G and the bond–bending graph H—too equally atom features, bond features and bail bending features—are taken as the inputs of GeoGNN.

a, In atom–bond graph G, the chemical bonds are regarded equally edges, connecting the atoms. In the bail–angle graph H, the bail angles are regarded as edges, and a bond angle connects two chemical bonds and three atoms. The double-dash arcs indicate the correspondence between the elements in the two graphs. b, Demonstration of geometry-level self-supervised learning tasks. The black circle represents the selected cantlet, whereas the greyness circles in graph Grand stand for the neighbouring masked atoms, the grey lines in graph G and the grey ovals in graph H represent the neighbouring masked bonds, and the grey lines in graph H stand for the neighbouring masked bond angles.
GeoGNN learns the representation vectors of atoms and bonds iteratively. For the yardth iteration, the representation vectors of cantlet u and bail (u,five) are denoted by h u and h u v , respectively. To connect the atom–bond graph G and bond–angle graph H, the representation vectors of the bonds are taken equally the advice links betwixt Grand and H. In the get-go pace, the bonds' representation vectors are learned by aggregating messages from the neighbouring bonds and corresponding bond angles in the bail–angle graph H. In the second step, the atoms' representation vectors are learned by aggregating messages from the neighbouring atoms and the corresponding bonds in the cantlet–bond graph M. Finally, the molecular representation h G is obtained by pooling over the atoms' representations. Encounter the Methods for details on the GeoGNN architecture.
Geometry-level self-supervised learning tasks
To farther boost the generalization power of GeoGNN, we suggest three geometry-level self-supervised learning tasks to pre-railroad train GeoGNN: (1) the bond lengths prediction; (2) the bond angles prediction; (3) the atomic distance matrices prediction. The bond lengths and bail angles describe the local spatial structures, whereas the atomic altitude matrices describe the global spatial structures.
Local spatial structures
Bond lengths and angles are the almost important molecular geometrical parameters: the former is the distance between two joint atoms in a molecule, reflecting the bond forcefulness between the atoms, whereas the latter is the angle connecting two consecutive bonds, including three atoms, describing the local spatial construction of a molecule.
To acquire the local spatial structures, we construct self-supervised learning tasks that predict bond lengths and angles. First, for a molecule, we randomly select 15% of atoms. For each selected atom, nosotros extract the ane-hop neighbourhood of this atom, including the side by side atoms and bonds, as well equally the bond angles formed by that selected atom. Second, we mask the features of these atoms, bonds and bond angles in the one-hop neighbourhood. The representation vectors of the extracted atoms and bonds at the final iteration of GeoGNN are used to predict the extracted bond lengths and bond angles. Cocky-supervised learning tasks based on bond lengths and bond angles are shown on left and middle of Fig. 2b. We design a regression loss function that penalizes the error between the predicted bond lengths/angles and the labels, whose details can be referred to in the Methods. The task of predicting the local spatial structures can exist seen as a node-level cocky-supervised learning task.
Global spatial structures
Except for the tasks for learning local spatial structures, we too design the atomic distance matrices prediction task for learning the global molecular geometry. We construct the atomic distance matrix for each molecule based on the three-dimensional coordinates of the atoms. Nosotros so predict the elements in the distance matrix, shown on the right of Fig. 2b.
Note that for 2 molecules with the same topological structures, the spatial distances betwixt the respective atoms could vary greatly; thus, for a molecule, rather than take predicting atomic altitude matrix equally a regression problem, we have it as a multi-class classification problem by projecting the diminutive distances into thirty bins with equal stride. Details on the designed loss function tin can exist institute in the Methods. The job predicting the bond lengths tin exist seen every bit a special example of the task predicting the diminutive distances. The old focuses more on the local spatial structures, whereas the latter focuses more on the distribution of the global spatial structures.
To pre-train GeoGNN, nosotros consider both the local spatial structures and global spatial structures for each molecule past summing up the corresponding loss functions.
Experiments
To thoroughly evaluate the performance of Gem, we compare it with multiple SOTA methods on multiple criterion datasets from MoleculeNet21 with diverse molecular property prediction tasks, such as concrete, chemical and biophysics.
Pre-grooming settings
Datasets
We use twenty million unlabelled molecules sampled from Zinc1522, a public access database that contains purchasable drug-similar compounds, to pre-train GeoGNN. Nosotros randomly sample 90% of the molecules for training and the remaining for evaluation.
Self-supervised learning chore settings
We utilize geometry- and graph-level tasks to pre-train GeoGNN. For the erstwhile, we utilise the Merck molecular force field (MMFF94)23 function in RDKit to obtain the simulated three-dimensional coordinates of the atoms in the molecules. The geometric features of the molecule—including bond lengths, bond angles and atomic distance matrices—are calculated by the false 3-dimensional coordinates. We predict the molecular fingerprints for the graph-level tasks. The graph-level tasks can exist formulated as a set of binary classification problems, where each bit of the fingerprints corresponds to one binary nomenclature problem. Two kinds of fingerprints are used: (i) the molecular access arrangement (MACCS) key24 and (2) the extended-connectivity fingerprint (ECFP)25.
Molecular property prediction settings
Datasets and splitting method
We conduct experiments on multiple molecular benchmarks from the MoleculeNet21, including both classification and regression tasks26,27,28,29,30,31. Post-obit the previous workeleven, nosotros split all the datasets with scaffold dissever32, which splits molecules according to the their scaffold (molecular substructure). Scaffold split is a more challenging splitting method and tin better evaluate the generalization ability of the models on out-of-distribution data samples.
GNN architecture
Nosotros use the Aggregate and COMBINE functions divers in the graph isomorphism network (GIN)19. Residual connections33, layer normalization34 and graph normalization35 are incorporated into GIN to further improve the performance. We also use the boilerplate pooling as the READOUT role to obtain the graph representation.
Evaluation metrics
Every bit suggested by the MoleculeNet21, we apply the boilerplate ROC-AUC36 as the evaluation metric for the classification datasets. ROC-AUC (area under the receiver operating feature curve) is used to evaluate the functioning of binary nomenclature tasks, for which higher is ameliorate. With respect to the regression datasets, we use root mean foursquare error (RMSE) for FreeSolv37, ESOL38 and Lipo39, whereas we utilize hateful average error (MAE) for QM740, QM841 and QM942. We execute iv independent runs for each method and report the mean and the standard deviation of the metrics.
Baselines
We compare the proposed method with various competitive baselines. D-MPNN43, AttentiveFP44, SGCNsixteen, DimeNet17 and HMGNN6 are the GNNs without pre-training, amidst which, SGCN, DimeNet and HMGNN contain three-dimensional geometry data; N-Gram45, PretrainGNN11 and GROVERfour are the methods with pre-training. N-Gram assembles the node embeddings in short walks in the graph and then leverages Random Forest or XGBoost to predict the molecular backdrop. PretrainGNN implements several types of self-supervised learning tasks, among which we report the best event. GROVER integrates GNN into Transformer with 2 self-supervised tasks, and we report the results of GROVERbase and GROVERbig with different network capacity.
Experimental results
Overall operation
The overall functioning of GEM along with other methods is summarized in Tabular array 1. We have the following observations: (1) Jewel achieves SOTA results on 14 out of xv datasets. On the regression tasks, Gem achieves an overall relative improvement of 8.8% on average compared with the previous SOTA results in each dataset. On the nomenclature tasks, Jewel achieves an overall relative comeback of 4.7% on the boilerplate ROC-AUC compared with the previous SOTA upshot from D-MPNN. (two) Jewel achieves more substantial improvements on the regression datasets than the classification datasets. Nosotros guess that the regression datasets focus on predicting the quantum chemic properties, which are highly correlated to molecular geometries.
Contribution of GeoGNN
We investigate the issue of GeoGNN without pre-training on the regression datasets, including the properties of quantum mechanics and physical chemical science, which are highly correlated to molecular geometries. GeoGNN is compared with multiple GNN architectures, including: (ane) the commonly used GNN architectures, GIN19, GAT46 and GCN47; (2) contempo works incorporating three-dimensional molecular geometry, SGCN16, DimeNet17 and HMGNNvi; (3) the architectures specially designed for molecular representation, D-MPNN43, AttentiveFP44 and GTransformer4. From Table 2, we can conclude that GeoGNN considerably outperforms other GNN architectures on all the regression datasets since GeoGNN incorporates geometrical parameters even though the three-dimensional coordinates of the atoms are faux. The overall relative improvement is 7.9% compared with the best results of previous methods.
Contribution of geometry-level tasks
To study the effect of the proposed geometry-level self-supervised learning tasks, we apply dissimilar types of self-supervised learning tasks to pre-train GeoGNN on the regression datasets. In Table 3, 'Without pre-railroad train' denotes the GeoGNN network without pre-training, 'Geometry' denotes our proposed geometry-level tasks, 'Graph' denotes the graph-level task that predicts the molecular fingerprints and 'Context'4 denotes a node-level job that predicts the diminutive context. In general, the methods with geometry-level tasks are meliorate than that without it. Furthermore, 'Geometry' performs better than 'Geometry + Graph' in the regression tasks, which may due to the weak connexion between molecular fingerprints and the regression tasks.
Pre-trained representations visualization
To intuitively observe the representations that the self-supervised tasks (without downstream fine-tuning) accept learned, we visualize the representations by mapping them to the 2-dimensional infinite by t-SNE algorithm48, whose details can be found in the Supplementary Information. The Davies Bouldin index49 is calculated to mensurate the separation of clusters. The lower the Davies Bouldin index, the better the separation of the clusters. Here we test whether the pre-training methods are able to distinguish molecules with valid geometry (generated from RDKit) from molecules with invalid geometry (random generated). Nosotros randomly select one,000 molecules from ZINC. For each molecule, we generate the valid and invalid geometry. Every bit shown in Fig. 3a, both the graph-level and geometry-level pre-preparation methods can better distinguish the valid geometry from invalid geometry compared to not pre-trained. Besides, the geometry-level pre-training can further subtract the Davies Bouldin Index to ii.63, compared with seven.88 of the graph-level.
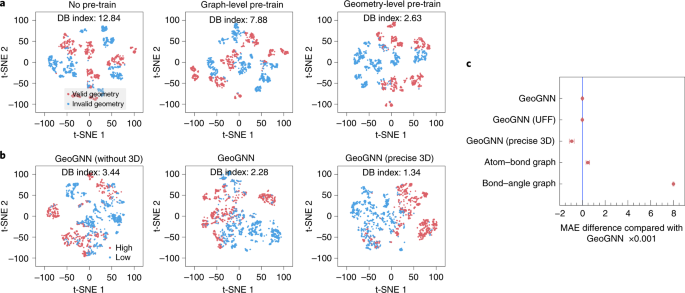
a, A pre-trained representation visualization comparing unlike self-supervised methods. The valid geometry cluster contains molecules with geometry generated by RDKit, whereas the invalid geometry cluster contains those with randomly generated geometry. b, A representation visualization comparing unlike qualities of geometries. The high cluster contains molecules with high Human–LUMO gaps, whereas the low cluster contains those with low Human being–LUMO gaps. c, MAE difference (the lower the better) on QM9 between baseline GeoGNN with other GeoGNN variants.
Impact of the quality of geometry
To investigate the impact of the quality of geometry, we commencement compare GeoGNN, which adopts the default force field MMFF, with GeoGNN (UFF), which adopts the universal force field (UFF)50, on dataset QM9. GeoGNN and GeoGNN (UFF) accomplish similar operation, as shown in Fig. 3c. The touch on of more than precise 3-dimensional coordinates provided past dataset QM9 (calculated by DFT51) is also investigated. GeoGNN (precise 3D) achieves a nifty comeback of about 12% compared with the baseline GeoGNN.
Furthermore, Fig. 3b shows the representation visuals for unlike qualities of molecular geometry. GeoGNN (without 3D) is a variant of GeoGNN that masks all the geometry features with zeros, GeoGNN is the baseline that utilizes coarse three-dimensional coordinates, and GeoGNN (precise 3D) utilizes precise 3D coordinates generated past DFT. Nosotros equally divide 2,000 molecules from QM9 into two clusters, 1 with loftier Homo–LUMO gaps and the other with low Human–LUMO gaps. We test the ability of dissimilar models to distinguish these two group of molecules. Visually, we observe that GeoGNN can better separate the clusters than GeoGNN (without 3D), whereas GeoGNN (precise 3D) works improve than GeoGNN. The differences in Davies Bouldin index support the observations.
Contributions of atom–bail and bond–bending graphs
We evaluate the contributions of the atom–bond and bail–bending graphs in GeoGNN on dataset QM9, as shown in Fig. 3c. Atom–bond graph utilizes the atom–bond graph just and pool over the representations of the atoms to judge the properties, whereas bond–angle graph utilizes the bond–angle graph only and pools over the representations of bonds. GeoGNN, which consists of both the atom–bond and bond–angle graphs, performs better than the above two variants, indicating that both the atom–bond and bail–angle graphs contribute to the performance.
Related work
Molecular representation
Current molecular representations can exist categorized into three types: molecular fingerprints, sequence-based representations and graph-based representations.
Molecular fingerprints
Molecular fingerprints such as ECFP25 and MACCS24 are molecular descriptors. Fingerprints are handcrafted representations—widely used by traditional machine learning methods3,52,53,54—that encode a molecule into a sequence of bits according to the molecules' topological substructures. Although fingerprints tin can represent the presence of the substructures in the molecules, they suffer from scrap collisions and vector sparsity, limiting their representation power.
Sequence-based representations
Some studies3,55 take SMILES strings56 that describe the molecules by strings as inputs, and leverage sequence-based models such as Recurrent Neural Networks and Transformer57,58 to learn the molecular representations; however, it is difficult for sequence-based methods to encompass the syntax of SMILES. For example, 2 adjacent atoms may be far apart in the text sequence. Likewise, a small change in a SMILES string tin lead to a large alter in the molecular structure.
Graph-based representations
Many works3,4,5,6,18 have showcased the great potential of graph neural networks on modelling molecules by taking each atom as a node and each chemical bond equally an edge. For example, AttentiveFP44 proposes to extend graph attending mechanism to learn assemblage weights. Meanwhile, a group of studies take tried to incorporate iii-dimensional geometry information: (1)13,fourteen,15 accept fractional geometry information equally features, such as atomic distances; (ii)sixteen proposed a spatial graph convolution that uses relative position vectors betwixt atoms as input features; (3)17 proposed a bulletin passing scheme based on bonds and transform letters from angles.
Pre-training for GNNs
Self-supervised learning7,viii,9,ten,59 has achieved great success in natural linguistic communication processing, figurer vision and other domains; it trains unlabelled samples in a supervised way to convalesce the overfitting issue and improve information utilization efficiency. Some studiesfour,eleven recently practical cocky-supervised learning methods to GNNs for molecular belongings prediction to overcome the insufficiency of the labelled samples. These works learn the molecular representation vectors past exploiting the node- and graph-level tasks, where the node-level tasks learn the local domain noesis past predicting the node backdrop and the graph-level tasks larn the global domain noesis by predicting biological activities. Although existing cocky-supervised learning methods can heave the generalization ability, they fail the spatial noesis that is strongly related to the molecular properties.
Decision
Efficient molecular representation learning is crucial for molecular property prediction. Existing works that use pre-training methods for molecular property prediction fail to utilize the molecular geometries described past bonds, bond angles and other geometrical parameters. To this end, nosotros pattern a geometry-based GNN and multiple geometry-level self-supervised learning methods capture the molecular spatial knowledge. Extensive experiments were conducted to verify the effectiveness of Gem, comparing it with multiple competitive baselines. Precious stone considerably outperforms other methods on multiple benchmarks. In the hereafter nosotros volition try to prefer the proposed framework to more molecular tasks, particularly the poly peptide–ligand affinity prediction job that requires lots of three-dimensional samplings.
Methods
Preliminary for GNNs
Graph neural networks is a message passing neural networks. More concretely, given a node 5, its representation vector \({\mathbf{h}}_{v}^{(m)}\) at the kth iteration is formalized past
$$\begin{array}{lll}{\mathbf{a}}_{v}^{(g)}&=&{\mathrm{Aggregate}}^{(k)}\left(\{({\mathbf{h}}_{5}^{(thou-i)},{\mathbf{h}}_{u}^{(k-ane)},{\mathbf{x}}_{uv}| u\in {{{\mathcal{Due north}}}}(v))\}\right.,\\ {\mathbf{h}}_{v}^{(yard)}&=&{\mathrm{COMBINE}}^{(g)}({\mathbf{h}}_{v}^{(k-ane)},{\mathbf{a}}_{v}^{(m)}).\end{array}$$
(1)
where \({{{\mathcal{Due north}}}}(five)\) is the ready of neighbours of node 5, AGGREGATE(k) is the aggregation function for accumulation messages from a node's neighbourhood, and COMBINE(thousand) is the update office for updating the node representation. We initialize \({\mathbf{h}}_{v}^{(0)}\) by the characteristic vector of node five, that is, \({\mathbf{h}}_{v}^{(0)}={\mathbf{ten}}_{v}\).
READOUT office is introduced to integrate the nodes' representation vectors at the final iteration so as to gain the graph'south representation vector h Yard , which is formalized as
$${\mathbf{h}}_{Yard}={\mathrm{READOUT}}({\mathbf{h}}_{v}^{(Chiliad)}| five\in {{{\mathcal{Five}}}}),$$
(2)
where K is the number of iterations. In virtually cases, READOUT is a permutation invariant pooling office, such as summation and maximization. The graph's representation vector h 1000 can then be used for downstream task predictions.
GeoGNN
The GeoGNN architecture encodes the molecular geometries by modelling two graphs: the cantlet–bond and bond–bending graphs, under which the representation vectors of atoms and bonds are learned iteratively. More concretely, the representation vectors of atom u and bond (u,five) for the kth iteration are denoted by h u and h u v , respectively. Nosotros initialize \({\mathbf{h}}_{u}^{(0)}={\mathbf{x}}_{u}\) and \({\mathbf{h}}_{uv}^{(0)}={\mathbf{ten}}_{uv}\).
Given bond (u,v), its representation vector \({\mathbf{h}}_{uv}^{(k)}\) at the kth iteration is formalized by
$$\begin{array}{ll}{\mathbf{a}}_{{uv}}^{(grand)}=&{\mathrm{AGGREGATE}}_{\mathrm{bail-angle}}^{(one thousand)}\left(\{({\mathbf{h}}_{uv}^{(thousand-1)},{\mathbf{h}}_{uw}^{(k-one)},{\mathbf{ten}}_{wuv}):westward\in {{{\mathcal{Northward}}}}(u)\}\right.\\ &\hspace{1em}\qquad \qquad \qquad \qquad \quad \left.\loving cup \{({\mathbf{h}}_{uv}^{(k-1)},{\mathbf{h}}_{vw}^{(k-one)},{\mathbf{x}}_{uvw}):w\in {{{\mathcal{N}}}}(v)\}\right),\\ {\mathbf{h}}_{uv}^{(k)}=&{\mathrm{COMBINE}}_{\mathrm{bond-angle}}^{(k)}({\mathbf{h}}_{uv}^{(k-1)},{\mathbf{a}}_{uv}^{(k)}).\terminate{array}$$
(3)
Hither, \({{{\mathcal{N}}}}(u)\) and \({{{\mathcal{North}}}}(v)\) denote the neighbouring atoms of u and v, respectively; \(\{(u,w):w\in {{{\mathcal{North}}}}(u)\}\cup \{(v,w):w\in {{{\mathcal{N}}}}(v)\}\) are the neighbouring bonds of (u,v). Amassbail−angle is the message aggregation function and COMBINEbail−angle is the update function for bond–angle graph H. In this manner, the information from the neighbouring bonds and the respective bail angles is aggregated into \({\mathbf{a}}_{uv}^{(k)}\). The representation vector of bond (u,five) is then updated according to the aggregated information. With the learned representation vectors of the bonds from bond–bending graph \({{{\mathcal{H}}}}\), given an atom u, its representation vector \({\mathbf{h}}_{u}^{(1000)}\) at the grandth iteration can be formalized equally
$$\begin{array}{lll}{\mathbf{a}}_{u}^{(thou)}&=&{\mathrm{AGGREGATE}}_{\mathrm{cantlet}-{\mathrm{bond}}}^{(k)}(\{({\mathbf{h}}_{u}^{(thou-one)},{\mathbf{h}}_{5}^{(g-one)},{\mathbf{h}}_{uv}^{(k-ane)}):v\in {{{\mathcal{N}}}}(u)\}),\\ {\mathbf{h}}_{u}^{(g)}&=&{\mathrm{COMBINE}}_{\mathrm{{atom}-{bond}}}^{(k)}({\mathbf{h}}_{u}^{(k-1)},{\mathbf{a}}_{u}^{(thou)}).\finish{assortment}$$
(4)
Similarly, \({{{\mathcal{Due north}}}}(u)\) denotes the neighbouring atoms of atom u, AGGREGATEatom−bond is the bulletin aggregation function for atom–bail graph G, and COMBINEatom−bond is the update function. For atom u, messages are aggregated from the neighbouring atoms and the corresponding bonds. Note that, the letters of the bonds are learned from the bond–angle graph H. The aggregated messages then update the representation vector of atom u.
The representation vectors of the atoms at the final iteration are integrated to proceeds the molecular representation vector h G by the READOUT part, which is formalized equally
$${\mathbf{h}}_{M}={\mathrm{READOUT}}({\mathbf{h}}_{u}^{(K)}| u\in {{{\mathcal{Five}}}}),$$
(5)
where K is the number of iterations. The molecule'south representation vector h G is used to predict the molecular properties.
Geometry-level cocky-supervised learning tasks
Local spatial structures
The cocky-supervised tasks for local spatial information are designed to learn 2 important molecular geometrical parameters, the bond lengths and the bond angles. The loss functions of the self-supervised tasks are defined as follows:
$$\brainstorm{array}{lll}{L}_{\mathrm{length}}({{{\mathcal{Due east}}}})&=&\frac{one}{| {{{\mathcal{E}}}}| }\mathop{\sum}\limits_{(u,five)\in {{{\mathcal{Due east}}}}}{({f}_{\mathrm{length}}({\mathbf{h}}_{u}^{(K)},{\mathbf{h}}_{v}^{(1000)})-{fifty}_{uv})}^{2};\\ {L}_{\mathrm{angle}}({{{\mathcal{A}}}})&=&\frac{1}{| {{{\mathcal{A}}}}| }\mathop{\sum}\limits_{(u,v,w)\in {{{\mathcal{A}}}}}{({f}_{\mathrm{bending}}({\mathbf{h}}_{u}^{(M)},{\mathbf{h}}_{v}^{(K)},{\mathbf{h}}_{westward}^{(One thousand)})-{\phi }_{uvw})}^{2}.\finish{array}$$
(6)
Here, \({Fifty}_{\mathrm{length}}({{{\mathcal{E}}}})\) is the loss function for bail lengths, with \({{{\mathcal{Eastward}}}}\) as the set of bonds; \({Fifty}_{\mathrm{angle}}({{{\mathcal{A}}}})\) is the loss part of bond angles, with \({{{\mathcal{A}}}}\) as gear up of angles; K is the number of iterations for GeoGNN; f length(⋅) is the network predicting the bond lengths; and f angle(⋅) is the network predicting the bond angles; l u v denotes the length of the bail connecting atoms u and 5; ϕ u v due west denotes the degree of the bond angle connecting bonds (u,v) and (v,w).
Global spatial structures
The cocky-supervised tasks for global spatial information are designed to learn the atomic altitude matrices between all atom pairs. Each element of the distance matrices is the 3-dimensional distance between two atoms. We use d u v to announce the distance between 2 atoms u and v in the molecule. For the atomic altitude prediction job, we clip the distance with the range from 0 Å to twenty Å and project it into 30 bins with equal stride. The loss function of the cocky-supervised tasks is defined equally follows:
$${L}_{\mathrm{distance}}({{{\mathcal{5}}}})=\frac{one}{| {{{\mathcal{V}}}}{| }^{ii}}\mathop{\sum}\limits_{u,five\in {{{\mathcal{Five}}}}}-{\mathrm{bin}}^{T}({d}_{uv})\cdot {\mathrm{log}}({f}_{\mathrm{distance}}({\mathbf{h}}_{u}^{(One thousand)},{\mathbf{h}}_{five}^{(Chiliad)})),$$
(7)
where \({{{\mathcal{V}}}}\) is the set of atoms, f altitude(⋅) is the network predicting the distribution of atomic distances, the bin(⋅) function is used to discretize the diminutive distance d u v into a one-hot vector and \({\mathrm{log}}(\cdot )\) is the logarithmic function.
Data availability
The self-supervised data used in our study are publicly available in ZINC (https://zinc.docking.org/tranches/dwelling/), whereas the downstream benchmarks tin be downloaded from MoleculeNet (https://moleculenet.org/datasets-1).
Code availability
The source code of this written report providing the geometry-based GNN and several geometry-level cocky-supervised learning methods is freely available at GitHub (https://github.com/PaddlePaddle/PaddleHelix/tree/dev/apps/pretrained_compound/ChemRL/Precious stone) to allow replication of the results. The version used for this publication is bachelor at https://doi.org/ten.5281/zenodo.5781821.
References
-
Shen, J. & Nicolaou, C. A. Molecular belongings prediction: recent trends in the era of bogus intelligence. Drug Discov. Today Technol. 32–33, 29–36 (2020).
-
Wieder, O. et al. A meaty review of molecular property prediction with graph neural networks. Drug Discov. Today Technol. 37, ane–12 (2020).
-
Huang, K. et al. DeepPurpose: a deep learning library for drug-target interaction prediction. Bioinformatics 36, 5545–5547 (2020).
-
Rong, Y. et al. Cocky-supervised graph transformer on big-calibration molecular data. In Advances in Neural Information Processing Systems 33: Annual Conference on Neural Data Processing Systems 2020 (eds Larochelle, H. et al.) 12559–12571 (NeurIPS 2020).
-
Shindo, H. & Matsumoto, Y. Gated graph recursive neural networks for molecular property prediction. Preprint at https://arxiv.org/abs/1909.00259 (2019).
-
Shui, Z. & Karypis, Thousand. Heterogeneous molecular graph neural networks for predicting molecule properties. In 20th IEEE International Conference on Information Mining (eds Establish, C. et al.) 492–500 (IEEE, 2020).
-
Devlin, J., Chang, M.-West., Lee, Grand. & Toutanova, Yard. BERT: pre-training of deep bidirectional transformers for linguistic communication agreement. In Proc. 2022 Conference of the North American Affiliate of the Association for Computational Linguistics: Human Language Technologies (eds Burstein, J. et al.) 4171–4186 (Association for Computational Linguistics, 2019).
-
He, P., Liu, X., Gao, J. & Chen, W. DeBERTa: decoding-enhanced BERT with disentangled attention. In ninth International Conference on Learning Representations (ICLR, 2021).
-
Doersch, C., Gupta, A. & Efros, A. A. Unsupervised visual representation learning by context prediction. In International Conference on Computer Vision (IEEE Computer Lodge, 2015).
-
Gidaris, S., Singh, P. & Komodakis, N. Unsupervised representation learning by predicting prototype rotations. In 6th International Conference on Learning Representations (ICLR, 2018).
-
Hu, W. et al. Strategies for pre-grooming graph neural networks. In eighth International Conference on Learning Representations (ICLR, 2020).
-
Peleg-Shulman, T., Najajreh, Y. & Gibson, D. Interactions of cisplatin and transplatin with proteins: comparing of binding kinetics, binding sites and reactivity of the pt-protein adducts of cisplatin and transplatin towards biological nucleophiles. J. Inorg. Biochem. 91, 306–311 (2002).
-
Schütt, K. et al. Schnet: A continuous-filter convolutional neural network for modeling quantum interactions. In Advances in Neural Data Processing Systems xxx: Annual Conference on Neural Information Processing Systems 2017 (eds Guyon, I. et al.) 991–1001 (NeurIPS, 2017).
-
Li, J., Xu, K., Chen, L., Zheng, Z. & Liu, Ten. GraphGallery: a platform for fast benchmarking and easy development of graph neural networks based intelligent software. In 43rd IEEE/ACM International Briefing on Software Engineering: Companion Proceedings thirteen–16 (IEEE, 2021).
-
Maziarka, L. et al. Molecule attention transformer. Preprint at https://arxiv.org/abs/2002.08264 (2020).
-
Danel, Tomasz et al. Spatial graph convolutional networks. In Neural Data Processing—27th International Conference, ICONIP 2020 Vol. 1333 (eds Yang, H. et al.) 668–675 (Springer, 2020).
-
Klicpera, J., Groß, J. & Günnemann, S. Directional message passing for molecular graphs. In 8th International Conference on Learning Representations (ICLR, 2020).
-
Gilmer, J., Schoenholz, S. S., Riley, P. F., Vinyals, O. & Dahl, G. E. Neural message passing for quantum chemical science. In Proc. 34th International Conference on Automobile Learning Vol. seventy (eds Precup, D. & Teh, Y. Due west.) 1263–1272 (PMLR, 2017).
-
Xu, Grand., Hu, W., Leskovec, J. & Jegelka, S. How powerful are graph neural networks? In 7th International Conference on Learning Representations (ICLR, 2019).
-
Lord's day, F.-Y., Hoffmann, J., Verma, Five. & Tang, J. Infograph: unsupervised and semi-supervised graph-level representation learning via mutual information maximization. In 8th International Briefing on Learning Representations (ICLR, 2020).
-
Wu, Z. et al. Moleculenet: a benchmark for molecular machine learning. Chem. Sci. 9, 513–530 (2018).
-
Sterling, T. & Irwin, J. J. ZINC 15—ligand discovery for everyone. J. Chem. Inf. Model. 55, 2324–2337 (2015).
-
Halgren, T. A. Merck molecular force field. I. Basis, grade, scope, parameterization, and performance of MMFF94. J. Comput. Chem. 17, 490–519 (1996).
-
Durant, J. 50., Leland, B. A., Henry, D. R. & Nourse, J. One thousand. Reoptimization of MDL keys for apply in drug discovery. J. Chem. Inf. Comput. Sci. 42, 1273–1280 (2002).
-
Rogers, D. & Hahn, M. Extended-connectivity fingerprints. J. Chem. Inf. Model. 50, 742–754 (2010).
-
Subramanian, G., Ramsundar, B., Pande, V. & Denny, R. A. Computational modeling of β-secretase i (bace-1) inhibitors using ligand based approaches. J. Chem. Inf. Model. 56, 1936–1949 (2016).
-
Martins, I. F., Teixeira, A. 50., Pinheiro, L. & Falcão, A. O. A Bayesian approach to in silico blood–brain barrier penetration modeling. J. Chem. Inf. Model. 52, 1686–1697 (2012).
-
Richard, A. M. et al. Toxcast chemical landscape: paving the road to 21st century toxicology. Chem. Res. Toxicol. 29, 1225–1251 (2016).
-
Gayvert, Yard. 1000., Madhukar, North. Due south. & Elemento, O. A data-driven approach to predicting successes and failures of clinical trials. Cell Chem. Biol. 23, 1294–1301 (2016).
-
Huang, R. et al. Editorial: Tox21 challenge to build predictive models of nuclear receptor and stress response pathways as mediated by exposure to ecology toxicants and drugs. Front. Environ. Sci. three, 85 (2017).
-
Kuhn, Yard., Letunic, I., Jensen, Fifty. J. & Bork, P. The SIDER database of drugs and side effects. Nucl. Acids Res. 44, 1075–1079 (2016).
-
Ramsundar, B., Eastman, P., Walters, P. & Pande, Five. Deep Learning for the Life Sciences: Applying Deep Learning to Genomics, Microscopy, Drug Discovery, and More (O'Reilly Media, 2019).
-
He, K., Zhang, X., Ren, S. & Sun, J. Deep rest learning for image recognition. In 2016 IEEE Briefing on Calculator Vision and Pattern Recognition 770–778 (IEEE, 2016).
-
Ba, L. J., Kiros, J. R. & Hinton, K. East. Layer normalization. In NIPS 2022 Deep Learning Symposium recommendation (NIPS, 2016).
-
Chen, Y., Tang, X., Qi, X., Li, C.-Yard. & Xiao, R. Learning graph normalization for graph neural networks. Preprint at https://arxiv.org/abs/2009.11746 (2020).
-
Bradley, A. P. The use of the area under the ROC bend in the evaluation of car learning algorithms. Blueprint Recognit. 30, 1145–1159 (1997).
-
Mobley, D. 50. & Guthrie, J. P. FreeSolv: a database of experimental and calculated hydration complimentary energies, with input files. J. Comput. Aided Mol. Des. 28, 711–720 (2014).
-
Delaney, J. S. ESOL: estimating aqueous solubility direct from molecular structure. J. Chem. Inf. Model. 44, 1000–1005 (2004).
-
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucl. Acids Res. twoscore, 1100–1107 (2012).
-
Blum, L. C. & Reymond, J.-Fifty. 970 Million druglike pocket-size molecules for virtual screening in the chemical universe database GDB-xiii. J. Am. Chem. Soc. 131, 8732–8733 (2009).
-
Ramakrishnan, R., Hartmann, M., Tapavicza, E. & AnatoleVonLilienfeld, O. Electronic spectra from TDDFT and machine learning in chemical infinite. J. Chem. Phys. 143, 084111 (2015).
-
Ruddigkeit, L., van Deursen, R., Blum, L. C. & Reymond, J.-50. Enumeration of 166 billion organic modest molecules in the chemical universe database GDB-17. J. Chem. Inf. Model. 52, 2864–2875 (2012).
-
Yang, K. et al. Analyzing learned molecular representations for belongings prediction. J. Chem. Inf. Model. 59, 3370–3388 (2019).
-
Xiong, Z. et al. Pushing the boundaries of molecular representation for drug discovery with the graph attending mechanism. J. Med. Chem. 63, 8749–8760 (2020).
-
Liu, S., Demirel, M. F. & Liang, Y. Northward-gram graph: simple unsupervised representation for graphs, with applications to molecules. In Advances in Neural Data Processing Systems 32: Almanac Conference on Neural Information Processing Systems 2019 (eds Wallach, H. One thousand. et al.) 8464–8476 (NeurIPS, 2019).
-
Velickovic, P. et al. Graph attention networks. In 5th International Conference on Learning Representations (ICLR, 2017).
-
Kipf, T. Due north. & Welling, M. Semi-supervised classification with graph convolutional networks. In fifth International Conference on Learning Representations (ICLR, 2017).
-
van der Maaten, L. Accelerating t-SNE using tree-based algorithms. J. Mach. Learn. Res. fifteen, 3221–3245 (2014).
-
Davies, D. L. & Bouldin, D. W. A cluster separation measure out. IEEE Trans. Blueprint Anal. Mach. Intell i, 224–227 (1979).
-
Rappé, A. K., Casewit, C. J., Colwell, K. S., Goddard, W. A. Three & Skiff, W. 1000. UFF, a total periodic tabular array force field for molecular mechanics and molecular dynamics simulations. J. Am. Chem. Soc. 114, 10024–10035 (1992).
-
Gross, E.K.U. & Dreizler, R. Grand. Density Functional Theory Vol. 337 (Springer, 2013).
-
Cereto-Massagué, A. et al. Molecular fingerprint similarity search in virtual screening. Methods 71, 58–63 (2015).
-
Coley, C. Westward., Barzilay, R., Green, W. H., Jaakkola, T. S. & Jensen, K. F. Convolutional embedding of attributed molecular graphs for physical property prediction. J. Chem. Inf. Model. 57, 1757–1772 (2017).
-
Duvenaud, D. et al. Convolutional networks on graphs for learning molecular fingerprints. In Advances in Neural Information Processing Systems 28: Annual Conference on Neural Data Processing Systems (eds Cortes, C. et al.) 2224–2232 (NeurIPS, 2015).
-
Goh, Chiliad. B., Hodas, N. O., Siegel, C. & Vishnu, A. SMILES2Vec: an interpretable full general-purpose deep neural network for predicting chemic properties. Preprint at https://arxiv.org/abs/1712.02034 (2018).
-
Weininger, D. SMILES, a chemical linguistic communication and information organisation. i. Introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28, 31–36 (1988).
-
Zaremba, Due west., Sutskever, I. & Vinyals, O. Recurrent neural network regularization. Preprint at https://arxiv.org/abs/1409.2329 (2014).
-
Vaswani, A. et al. Attention is all you need. In Advances in Neural Information Processing Systems thirty: Annual Conferenceon Neural Data Processing Systems 2017 5998–6008 (NeurIPS, 2017).
-
Li, P. et al. Learn molecular representations from big-scale unlabeled molecules for drug discovery. Preprint at https://arxiv.org/abs/2012.11175 (2020).
Acknowledgements
This work is supported past National Engineering Research Center of Deep Learning Engineering and Applications.
Author information
Affiliations
Contributions
X.F., F.W., H. Wu and H. Wang led the inquiry. L.L., 10.F. and F.Due west. contributed technical ideas. 50.L., J.L., D.H., S.Z. and X.F. adult the proposed method. 10.F., L.L., S.Z. and J.Z. developed analytics. X.F., L.L, F.Westward., J.L., D.H., Southward.Z. and J.Z. wrote the paper.
Corresponding authors
Ideals declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open up Admission This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, accommodation, distribution and reproduction in any medium or format, as long as you give advisable credit to the original writer(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other tertiary party material in this article are included in the article's Creative Commons license, unless indicated otherwise in a credit line to the textile. If material is non included in the article's Creative Eatables license and your intended use is non permitted by statutory regulation or exceeds the permitted use, yous volition demand to obtain permission directly from the copyright holder. To view a re-create of this license, visit http://creativecommons.org/licenses/by/4.0/.
Reprints and Permissions
About this article
Cite this article
Fang, X., Liu, L., Lei, J. et al. Geometry-enhanced molecular representation learning for belongings prediction. Nat Mach Intell 4, 127–134 (2022). https://doi.org/10.1038/s42256-021-00438-4
-
Received:
-
Accustomed:
-
Published:
-
Issue Date:
-
DOI : https://doi.org/x.1038/s42256-021-00438-4
stapletonlonot1975.blogspot.com
Source: https://www.nature.com/articles/s42256-021-00438-4
0 Response to "array of ai 3d drawing nature geometry"
แสดงความคิดเห็น